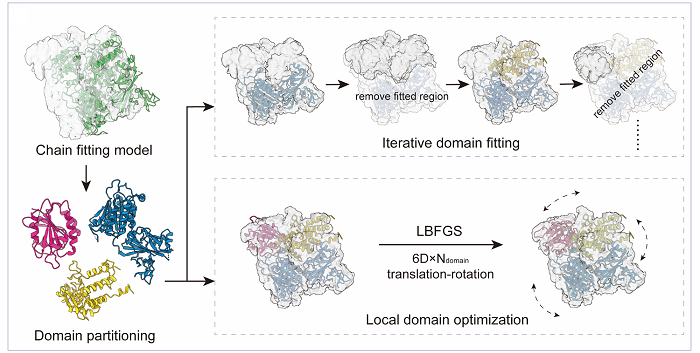
Domain-level assembly and optimization:
Domain-based fitting strategies. The chain is segmented into multiple domains. Proteins without homologous chains undergo iterative domain fitting. Each domain is individually matched to the density map in a long-to-short order based on sequence length, using the limited-memory Broyden-Fletcher-Goldfarb-Shanno (L-BFGS) algorithm, guided by the density correlation energy function.
For proteins with homologous protein chains, local domain optimization is performed using the L-BFGS algorithm to refine the position and orientation of each domain within the chain, guided by density correlation and physicochemical energy functions.